Universal-Flex Multiplex qPCR-Master Mix, with ROX-alike reference dye
The qPCR Multiplex Master mix offers quantitative PCR with a new ROX-alike reference dye for using with all common PCR-Platforms.
The Universal-Flex Multiplex qPCR Mastermix is compatible across a variety of instrument platforms including those that require a high, low or no ROX reference signal
Components and Mixture
The Universal-flex Multiplex qPCR Master Mix for probes (2X) is a reaction mix optimized for real-time qPCR detection and quantitation of target DNA up to four sequences using hydrolysis probes. It contains Hot Start Taq DNA polymerase and all components like dNTPs (dUTP instead dTTP), UDG, MgCL2, reaction buffer additives and stabilizers, optimized for Probe qPCR to ensures:
Features:
- a perfect amplification curve
- no need to adjust ROX concentration
- accurate quantification of target genes
- increased PCR specificity and sensitivity
- good repeatability and high reliability
- over a broad dynamic range
- UDG/dUTP prevent residual contamination
Applications:
- Real-time PCR with up to four target-sequences (valid for all ROX concentrations)
- Conventional PCR (cDNA, plasmid DNA, Lambda DNA)
- High-throughput PCR
Description:
The 2X Mastermix contains all components all reagents required for qPCR (except template and primers). At the same time, the product contains a special ROX Passive Reference Dye, which is suitable for use in all qPCR instruments, and there is no need to adjust the ROX concentration on different instruments.
Only templates, primers, probes and Nuclease-free Water need to be added for use. It also features a unique passive reference dye that is compatible across a variety of instrument platforms and a non-fluorescent blue visible dye to monitor reaction setup. This dye does not spectrally overlap fluorophores commonly used for qPCR and will not interfere with real-time detection.
Taqs: Luna, NEB, Universal Multiplex Mastermix for probes
REDUCED PRICE!
Final price excl. shipping costs3
- verfügbar / avaílable
- 1 - 3 Tage Lieferzeit / 1-3 days delivery1
Storage and
transportation: at -20 °С;. Shipping
with blue ice or at room temperature
Storage
terms: up to 18 months
Please note:
1. After thawing, please gently mix up and down, do not vortex, avoid bubbles, mix well before use.
2. When preparing the reaction solution, please place the reagent on the ice.
3. The product contains fluorescent dyes, so strong light should be avoided when preparing qPCR reaction solution.
4. New disposable tips should be used for preparation of reaction mixes to avoid cross contamination.
5. Avoid freeze-thawing cycles of the Master Mix, and try to use it up within a month after thawing.
The Universal-Flex Multiplex qPCR Mastermix is compatible across a variety of instrument platforms including those that require a high, low or no ROX reference signal
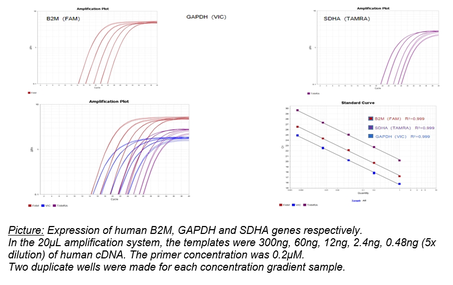
Picture: Expression of human B2M, GAPDH and SDHA genes respectively.
In the 20μL amplification system, the templates were 300ng, 60ng, 12ng, 2.4ng, 0.48ng (5x dilution) of human cDNA. The primer concentration was 0.2μM.
Two duplicate wells were made for each concentration gradient sample.
The experimental results as shown in the picture above, the lower left picture is the result of superposition of three genes, and the lower right picture is the standard song.
Universal qPCR Mastermix Amplification protocol
1. Defrost the reaction mixture and stir thoroughly.
2. Add the following components into the thin-wall PCR tubes considering the final volume of a reaction mixture equal to 20 μl:
| Component | Volume | Final concentration |
| 2x Mastermix | 10 µl | 1x |
| Each Forward Primer (10µM) *1 | variable | 0,1 - 0,5 µM |
|
Each Reverse Primer(10µM)*1 |
variable | 0,1 - 0,5 µM |
| Probe (10 µM) *1 | variable | 0,1 - 0,5 µM |
| DNA Template *2 | variable | 10 pg - 1 µg |
| Sterile Water | up to 20 µl |
3. Gently vortex and remove droplets by centrifugation.
4. Perform PCR
*1: a: Usually, a good amplification effect can be obtained with the final concentration of 0.2 μM. When the reaction performance is poor, the primer concentration can be adjusted in the range of 0.2-1.0 μM.
*2: b: The amount of template added varies depending on the number of copies of the target gene, and the appropriate amount of template addition is studied by gradient dilution. The best addition amount of template DNA in the 20 µl reaction system was less than 100 ng.
Universal qPCR Mastermix Cycler program
| Step |
Temp. °C |
Incubation time
|
Number of Cycles |
|
UDG incobation |
50 | 2 min | 1 |
|
Preliminary denaturation |
95 | 0,5-2 min | 1 |
|
Denaturation |
95 | 15 sec | 30-40 |
|
Annealing |
55-65 |
10 sec |
30-40 |
| Elongation | 72 | 30 sec | 30-40 *1 |
|
Melting curve (recommended) |
1 | 1 |
as an alternative:
| Step |
Temp. °C |
Incubation time
|
Number of Cycles |
|
UDG incubation |
50 | 2 min | 1 |
|
Preliminary denaturation |
95 | 0,5-2 min | 1 |
|
Denaturation |
95 | 15 sec | 30-40 |
|
Annealing / Extension |
60 |
30 sec |
30-40 *1 |
| Elongation | 72 | 30 sec | 30-40 |
|
Melting curve (recommended) |
1 | 1 |
*1: If amplification specificity needs to be improved, two-step procedure or annealing temperature can be used; To improve the amplification efficiency, a three-step procedure or extension time can be used.
Compatible instruments / Cycler List
ABI: 5700, 7000, 7300, 7700, 7900, 7900HT, 7900 HT Fast, StepOne™, StepOne Plus™, 7500/7500 Fast, ViiA 7™,;
Analytik Jena: qTOWER series;
qTOWER: LineGene series
Stratagene: Mx3000P®, 3005P™, 4000™;
Bio-Rad: CFX96™, CFX384™, iCycler iQ™, iQ5™, MyiQ™, MiniOpticon™, Opticon®, Opticon 2, Chromo4™;
Eppendorf: Realplex 2s, Mastercycler® ep, Realplex;
IIIumina: Eco QPCR;
Cepheid: SmartCycler®; QuantStudio™ series, PikoRealTM Cycler
Qiagen Corbett: Rotor-Gene® series;
Roche: LightCycler™ series;
Takara: Thermal Cycler Dice series;
Deutsche Beschreibung

Bio-Star Universal-Flex Multiplex qPCR-Mastermix with a novel ROX-likely reference dye
Bio-Star Universal-Flex qPCR-Mastermix with SYBRGREEN Blue and a novel ROX-likely reference dye
Material Safety Datasheet

References / Protocols / Notes / Recomendations / Tests
Primer Design Principles
Taqman primer design principle
1. Determine the probe before designing primers.
2. When designing primers, get as close to the probe as possible without overlapping the probe.
3. Avoid using 4 or more consecutive G.
4. The Tm value of each primer should be 58-60°C.
5. The last 5 nucleotides at the end of the primer cannot have more than 2 G and C.
6. Primers had better not contain self-complementary sequences, otherwise they will form the hairpins.
7. In order to avoid the amplification of the genome, it is best to design primers across exons.
8. The length of amplification product should be 50-150 bp in order to obtain the best PCR efficiency.
9. No other non-specific products were found in the comparison results on NCBI.
Taqman probe design principle
1. Probe length should be 13-25 bp (13-30 bp if conventional TaqMan probe is used).
2. The Tm value should be 65°C~70°C, which is usually 5°C~10°C higher than the TM value of the primer to ensure that the probe preferentially binds to the target gene during annealing.
3. For a primer, the content of guanine-cytosine (G+C) should be between 40% and 70%.
4. The 5 'end of the probe should avoid using G, because the 5' end G will have quenching effect, even if it is cut off.
5. In the whole probe, the content of C is obviously higher than that of G, and the high content of G will have quenching effect, so we can choose another paired chain as the probe.
Taqman MGB probe design principle
1. A report dye (for example, FAMTM) is attached to the 5 'end of the probe.
2. There is a non-fluorescence quenching group (NFQ) at the 3 'end of the probe.
3. The part of MGB is attached to NFQ, and MGBs increases the annealing temperature (Tm) without increasing the length of the probe, so a shorter probe can be designed, but not less than 13 bp.
4. In principle, as long as there is a base mutation in the MGB probe, MGB can detect it (the MGB probe will not bind to the target gene and will not produce a fluorescent signal).